Output in zarr (advanced)#
Topics:
Capturing output in memory using the
MemoryStoreofzarr(link)Transferring data from a
MemoryStoreinto a directory (viaDirectoryStore; link) or a zip file (viaZipStore; link).Re-chunking output to better align with specific analyses (time vs. spatial filtering).
A simple flow field#
We’ll use a Rankine Vortex (https://en.wikipedia.org/wiki/Rankine_vortex) and add some noise to make trajectories interesting.
[1]:
import matplotlib.pyplot as plt
import numpy as np
import xarray as xr
[2]:
np.random.seed(1234) # let's be reproducible
[3]:
lat = xr.DataArray(np.linspace(-20, 20, 101), dims=("lat",), name="lat")
lon = xr.DataArray(np.linspace(-20, 20, 101), dims=("lon",), name="lon")
# rankine vortex:
Vmax = 1.5
Vnoise = 0.2
phi = np.arctan2(lat, lon)
r = (lat**2 + lon**2) ** 0.5
core_rad = 10
rad_structure = xr.where(r <= core_rad, r / core_rad, core_rad / r)
U_noise = np.random.normal(0, Vnoise, size=r.shape)
V_noise = np.random.normal(0, Vnoise, size=r.shape)
# the dataset
ds_flow_field = xr.Dataset(
{
"U": (Vmax * -np.sin(phi) * rad_structure + U_noise).rename("U"),
"V": (Vmax * np.cos(phi) * rad_structure + V_noise).rename("V"),
},
coords={"lat": lat, "lon": lon},
)
display(ds_flow_field)
((ds_flow_field.U**2 + ds_flow_field.V**2) ** 0.5).plot()
(
ds_flow_field.isel(lon=slice(None, None, 3), lat=slice(None, None, 3)).plot.quiver(
x="lon", y="lat", u="U", v="V"
)
);
<xarray.Dataset>
Dimensions: (lat: 101, lon: 101)
Coordinates:
* lat (lat) float64 -20.0 -19.6 -19.2 -18.8 -18.4 ... 18.8 19.2 19.6 20.0
* lon (lon) float64 -20.0 -19.6 -19.2 -18.8 -18.4 ... 18.8 19.2 19.6 20.0
Data variables:
U (lat, lon) float64 0.4693 0.1444 0.6768 ... -0.2509 -0.1513 -0.425
V (lat, lon) float64 -0.4215 -0.3119 -0.2634 ... 0.02258 0.4363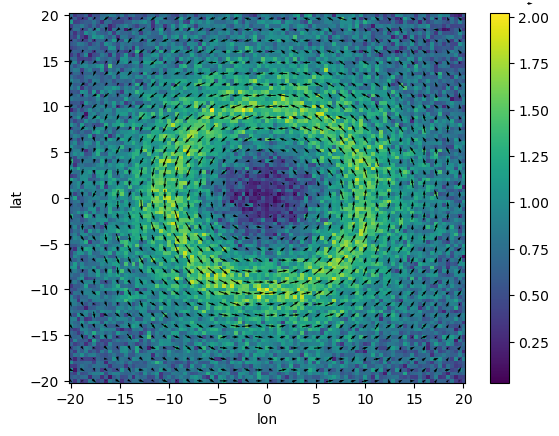
The Parcels experiment#
[4]:
from datetime import datetime, timedelta
import numpy as np
import zarr
from parcels import AdvectionRK4, FieldSet, ParticleSet, ScipyParticle
Create fieldset from Xarray#
[5]:
fieldset = FieldSet.from_xarray_dataset(
ds_flow_field,
variables={"U": "U", "V": "V"},
dimensions={"lat": "lat", "lon": "lon"},
)
A randomly positioned particleset#
[6]:
def create_random_pset(
fieldset=None, lon_range=(-15, 15), lat_range=(-15, 15), number_particles=200
):
return ParticleSet.from_list(
fieldset=fieldset,
pclass=ScipyParticle,
lon=np.random.uniform(*lon_range, size=(number_particles,)),
lat=np.random.uniform(*lat_range, size=(number_particles,)),
time=np.zeros(shape=(number_particles,)),
)
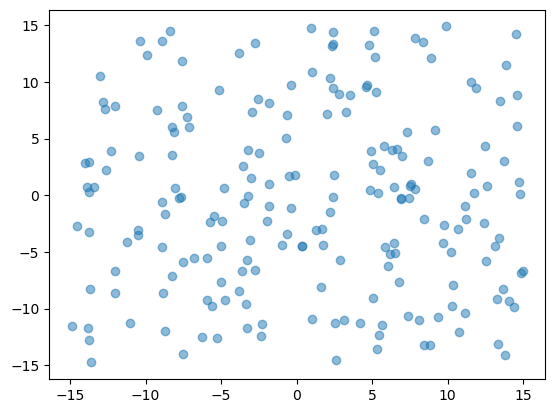
pset = create_random_pset(fieldset)
plt.plot(pset.lon, pset.lat, "o", alpha=0.5)
[6]:
[<matplotlib.lines.Line2D at 0x13f3d2c90>]
Streaming to an in-memory store#
[7]:
# create fresh particle set
pset = create_random_pset(fieldset)
# create output store and output file
output_memorystore = zarr.storage.MemoryStore()
outputfile = pset.ParticleFile(name=output_memorystore, outputdt=timedelta(hours=3))
# run experiment
pset.execute(
AdvectionRK4,
runtime=timedelta(days=17),
dt=timedelta(hours=3),
output_file=outputfile,
)
# load output
ds_out = xr.open_zarr(output_memorystore)
display(ds_out)
# have a look
ds_out.to_dataframe().plot.scatter(x="lon", y="lat", s=2, alpha=0.1);
INFO: Output files are stored in <zarr.storage.MemoryStore object at 0x164133550>.
100%|██████████| 1468800.0/1468800.0 [00:06<00:00, 240698.79it/s]
<xarray.Dataset>
Dimensions: (trajectory: 200, obs: 136)
Coordinates:
* obs (obs) int32 0 1 2 3 4 5 6 7 ... 128 129 130 131 132 133 134 135
* trajectory (trajectory) int64 200 201 202 203 204 ... 395 396 397 398 399
Data variables:
lat (trajectory, obs) float32 dask.array<chunksize=(200, 1), meta=np.ndarray>
lon (trajectory, obs) float32 dask.array<chunksize=(200, 1), meta=np.ndarray>
time (trajectory, obs) timedelta64[ns] dask.array<chunksize=(200, 1), meta=np.ndarray>
z (trajectory, obs) float32 dask.array<chunksize=(200, 1), meta=np.ndarray>
Attributes:
Conventions: CF-1.6/CF-1.7
feature_type: trajectory
ncei_template_version: NCEI_NetCDF_Trajectory_Template_v2.0
parcels_kernels: ScipyParticleAdvectionRK4
parcels_mesh: spherical
parcels_version: v2.4.2-370-gd0cb4110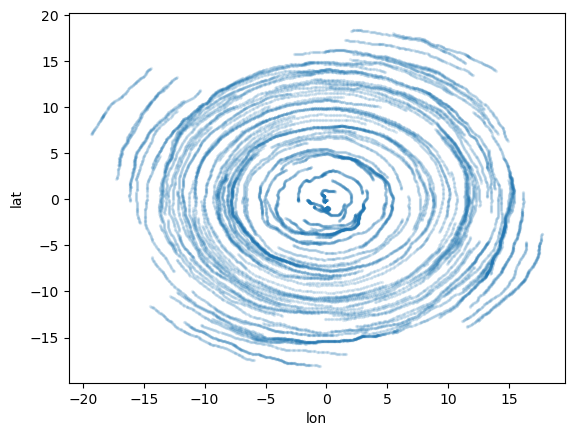
Saving to an other Zarr store#
Directory, without changing chunking#
[8]:
# create store
output_dirstore_name = "zarr_advanced_01.zarr/"
!rm -rf {output_dirstore_name}
output_dirstore = zarr.storage.DirectoryStore(output_dirstore_name)
# copy
zarr.convenience.copy_store(output_memorystore, output_dirstore)
output_dirstore.close()
# have a quick look
ds_out = xr.open_zarr(output_dirstore_name)
display(ds_out)
ds_out.to_dataframe().plot.scatter(x="lon", y="lat", s=2, alpha=0.1);
<xarray.Dataset>
Dimensions: (trajectory: 200, obs: 136)
Coordinates:
* obs (obs) int32 0 1 2 3 4 5 6 7 ... 128 129 130 131 132 133 134 135
* trajectory (trajectory) int64 200 201 202 203 204 ... 395 396 397 398 399
Data variables:
lat (trajectory, obs) float32 dask.array<chunksize=(200, 1), meta=np.ndarray>
lon (trajectory, obs) float32 dask.array<chunksize=(200, 1), meta=np.ndarray>
time (trajectory, obs) timedelta64[ns] dask.array<chunksize=(200, 1), meta=np.ndarray>
z (trajectory, obs) float32 dask.array<chunksize=(200, 1), meta=np.ndarray>
Attributes:
Conventions: CF-1.6/CF-1.7
feature_type: trajectory
ncei_template_version: NCEI_NetCDF_Trajectory_Template_v2.0
parcels_kernels: ScipyParticleAdvectionRK4
parcels_mesh: spherical
parcels_version: v2.4.2-370-gd0cb4110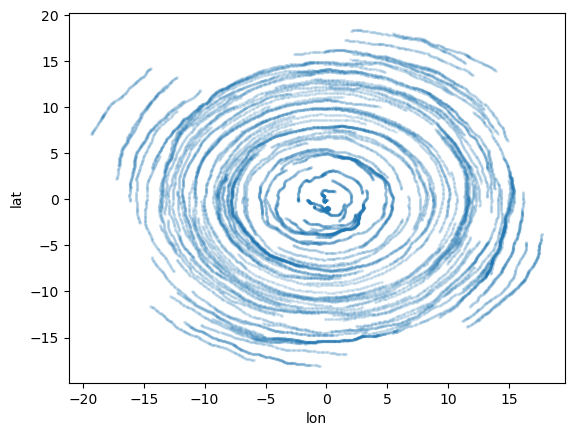
Zipfile, without changing chunking#
[9]:
# create store
output_zipstore_name = "zarr_advanced_01.zip"
!rm -f {output_zipstore_name}
output_zipstore = zarr.storage.ZipStore(output_zipstore_name, mode="w")
# copy
zarr.convenience.copy_store(output_memorystore, output_zipstore)
output_zipstore.close()
# have a quick look
ds_out = xr.open_zarr(output_zipstore_name)
display(ds_out)
ds_out.to_dataframe().plot.scatter(x="lon", y="lat", s=2, alpha=0.1);
<xarray.Dataset>
Dimensions: (trajectory: 200, obs: 136)
Coordinates:
* obs (obs) int32 0 1 2 3 4 5 6 7 ... 128 129 130 131 132 133 134 135
* trajectory (trajectory) int64 200 201 202 203 204 ... 395 396 397 398 399
Data variables:
lat (trajectory, obs) float32 dask.array<chunksize=(200, 1), meta=np.ndarray>
lon (trajectory, obs) float32 dask.array<chunksize=(200, 1), meta=np.ndarray>
time (trajectory, obs) timedelta64[ns] dask.array<chunksize=(200, 1), meta=np.ndarray>
z (trajectory, obs) float32 dask.array<chunksize=(200, 1), meta=np.ndarray>
Attributes:
Conventions: CF-1.6/CF-1.7
feature_type: trajectory
ncei_template_version: NCEI_NetCDF_Trajectory_Template_v2.0
parcels_kernels: ScipyParticleAdvectionRK4
parcels_mesh: spherical
parcels_version: v2.4.2-370-gd0cb4110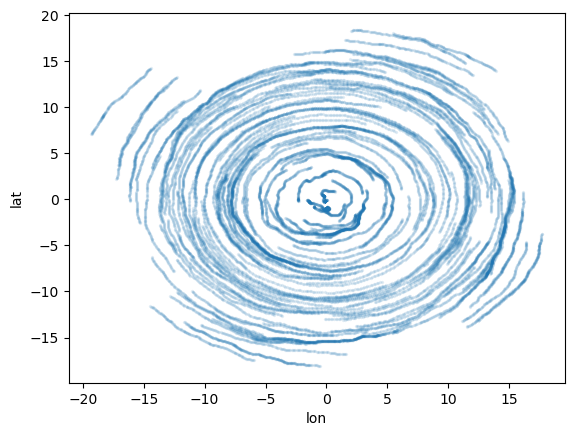
Rechunking and saving to new stores#
Consider the current chunking (let’s load the memory store again):
[10]:
ds_out_orig = xr.open_zarr(output_memorystore)
display(ds_out_orig.lat.data)
display(ds_out_orig.chunks)
|
||||||||||||||||
Frozen({'trajectory': (200,), 'obs': (1, 1, 1, 1, 1, 1, 1, 1, 1, 1, 1, 1, 1, 1, 1, 1, 1, 1, 1, 1, 1, 1, 1, 1, 1, 1, 1, 1, 1, 1, 1, 1, 1, 1, 1, 1, 1, 1, 1, 1, 1, 1, 1, 1, 1, 1, 1, 1, 1, 1, 1, 1, 1, 1, 1, 1, 1, 1, 1, 1, 1, 1, 1, 1, 1, 1, 1, 1, 1, 1, 1, 1, 1, 1, 1, 1, 1, 1, 1, 1, 1, 1, 1, 1, 1, 1, 1, 1, 1, 1, 1, 1, 1, 1, 1, 1, 1, 1, 1, 1, 1, 1, 1, 1, 1, 1, 1, 1, 1, 1, 1, 1, 1, 1, 1, 1, 1, 1, 1, 1, 1, 1, 1, 1, 1, 1, 1, 1, 1, 1, 1, 1, 1, 1, 1, 1)})
So we have one chunk per obs slice which covers all trajectories. This is the optimal way of creating the recorded dataset at simulation time, but may not be ideal for, e.g., time filtering.
So let’s turn this into a store which only covers 10 trajectories per chunk but 40 time steps:
[11]:
ds_out_rechunked = ds_out_orig.chunk({"trajectory": 10, "obs": 40})
display(ds_out_rechunked.lat.data)
display(ds_out_rechunked.chunks)
|
||||||||||||||||
Frozen({'trajectory': (10, 10, 10, 10, 10, 10, 10, 10, 10, 10, 10, 10, 10, 10, 10, 10, 10, 10, 10, 10), 'obs': (40, 40, 40, 16)})
And of course, we could now save the rechunked data into a zip store or a directory store:
[12]:
# necessary because of https://github.com/pydata/xarray/issues/4380
for vname, vobj in ds_out_rechunked.data_vars.items():
if "chunks" in vobj.encoding:
del vobj.encoding["chunks"]
[13]:
ds_out_rechunked.to_zarr("zarr_advanced_01_rechunked.zarr/", mode="w")
# new directory store
ds_out_rechunked.to_zarr(
zarr.storage.ZipStore("zarr_advanced_01_rechunked.zip", mode="w")
); # new zip store
